Combinatorial Proteomic®
Combinatorial Proteomic® uses antibody libraries as probes in order to profile the expression and function of proteins in complex proteomes.
The use of antibody libraries allows the detection of iper- and ipo- expressed proteins even at pico-quantity levels, overcoming one of the proteomic limitations: the difficulty in detecting low abundance proteins.
The goal of Combinatorial Proteomic® is the global analysis of protein expression and function.
Identifying proteins that are differentially expressed in specific cancers will
provide a wide range of novel biomarkers to be used in the creation of diagnostic and prognostic tools.
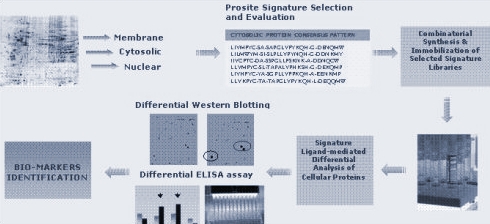
Combinatorial Proteomic® is a process that includes 5 main phases:
1. in silico analysis of human proteome using the pattern matcher PatScan and Prosite database
2. combinatorial synthesis of selected signature libraries by Fmoc chemistry and Mix & Split methods
3. immobilization on solid support of selected signature libraries
4. purification of signature-specific IgM antibodies by affinity chromatography experiments
5. differential analysis of cellular protein expression levels